ppALIGN API documentation
ProteinScoring Class Reference
#include <scorematrix.hpp>
Inheritance diagram for ProteinScoring:
Public Member Functions | |
| ProteinScoring (const std::string &name) | |
Static Public Member Functions | |
| static double | GetScale (const std::string &name) |
Detailed Description
Integer valued score matrix for proteins.
Definition at line 170 of file scorematrix.hpp.
Constructor & Destructor Documentation
| ProteinScoring::ProteinScoring | ( | const std::string & | name | ) |
Constructor.
- Parameters:
-
name matrix name (e.g. blosum62)
Member Function Documentation
| static double ProteinScoring::GetScale | ( | const std::string & | name | ) | [inline, static] |
Scale 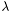 of the named score matrix. The scale is defined through:
of the named score matrix. The scale is defined through: ![$ s[a][b]= \lambda \log \frac{p(a,b)}{q(a)q(b)} $](form_10.png) .
.
- Parameters:
-
a1 alphabet of the first sequence a2 alphabet of the second sequence name name of the protein score matrix
Definition at line 186 of file scorematrix.hpp.
The documentation for this class was generated from the following file:
- src/scorematrix.hpp