ppALIGN API documentation
ScoreMatrix Class Reference
#include <scorematrix.hpp>
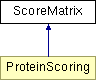
Public Member Functions | |
| ScoreMatrix (const Alphabet &a1, const Alphabet &a2) | |
| ScoreMatrix (const Alphabet &a1, const Alphabet &a2, int match, int mismatch) | |
| ScoreMatrix (const Alphabet &a1, const Alphabet &a2, const std::string &name) | |
| const Alphabet & | GetAlphabet1 () const |
| const Alphabet & | GetAlphabet2 () const |
| double | GetScale (const Frequencies &freq1, const Frequencies &freq2) const |
| double | GetScale (const Sequence &seq1, const Sequence &seq2) const |
Static Public Member Functions | |
| static double | GetScale (const Alphabet &a1, const Alphabet &a2, const std::string name) |
Detailed Description
Integer valued score matrix.
- Examples:
Definition at line 31 of file scorematrix.hpp.
Constructor & Destructor Documentation
Construct a score matrix.
- Parameters:
-
a1 alphabet for the first index a2 alphabet for the second index
Definition at line 41 of file scorematrix.hpp.
| ScoreMatrix::ScoreMatrix | ( | const Alphabet & | a1, | |
| const Alphabet & | a2, | |||
| int | match, | |||
| int | mismatch | |||
| ) | [inline] |
Construct a score matrix and initalize.
- Parameters:
-
a1 alphabet for the first index a2 alphabet for the second index match set diagonal (i==j) values to match mismatch set off-diagonal (i!=j) values to mismatch
Definition at line 62 of file scorematrix.hpp.
Construct a score matrix by name or by file. If name is not a valid matrix name, try to open a file and read the data from that.
Member Function Documentation
| const Alphabet& ScoreMatrix::GetAlphabet1 | ( | ) | const [inline] |
Alphabetbet.
- Returns:
- underlying alphabet of the first sequence.
Definition at line 88 of file scorematrix.hpp.
| const Alphabet& ScoreMatrix::GetAlphabet2 | ( | ) | const [inline] |
Alphabetbet.
- Returns:
- underlying alphabet of the second sequence.
Definition at line 96 of file scorematrix.hpp.
Estimate scale from input sequences. If &GetAlphabet1()==&GetAlphabet2() the background frequencies q(a) are estimated from both sequences and 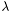 is determined as the root of
is determined as the root of
![\[ \sum_{a,b} q(a) q(b) e^{s[a][b])/\lambda} = 1 \]](form_16.png)
If &GetAlphabet1()!=&GetAlphabet2() the background frequencies are determined for both sequences separtely q_1(a) and q_2 and 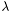 is determined as the root of
is determined as the root of
![\[ \sum_{a,b} q_1(a) q_2(b) e^{s[a][b])/\lambda} = 1 \]](form_17.png)
- Parameters:
-
seq1 seq2
| double ScoreMatrix::GetScale | ( | const Frequencies & | freq1, | |
| const Frequencies & | freq2 | |||
| ) | const |
Determine scale of the score matrix. The scale is defined through ![$ s[a][b]= \lambda \log \frac{p(a,b)}{q(a)q(b)} $](form_10.png) . It is estimated by the unique solution of the equation
. It is estimated by the unique solution of the equation
![\[ \sum_{a,b} freq1[a] freq2[b] e^{s[a][b])/\lambda} = 1 \]](form_15.png)
- Parameters:
-
freq1 background frequencies (assumed to be normalized) freq2 background frequencies (assumed to be normalized)
| static double ScoreMatrix::GetScale | ( | const Alphabet & | a1, | |
| const Alphabet & | a2, | |||
| const std::string | name | |||
| ) | [static] |
Scale 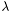 of the named score matrix. The scale is defined through:
of the named score matrix. The scale is defined through: ![$ s[a][b]= \lambda \log \frac{p(a,b)}{q(a)q(b)} $](form_10.png) .
.
The scales of the blosum matrices are:
- blosum30 : 1/5 bit
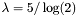
- blosum35-blosum40 : 1/4 bit,
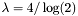
- blosum45-blosum55: 1/3 bit,
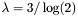
- blosum60-blosum100: 1/2 bit,
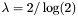
The scales of the pam matrices are:
- pam10-pam160 1/2 bit,
- pam170-pam260 1/3 bit,
- pam270-pam340 1/4 bit,
- pam350-pam400 1/5 bit,
- pam410-pam460 1/6 bit,
- pam470-pam500 1/7 bit
- Parameters:
-
a1 alphabet of the first sequence a2 alphabet of the second sequence name name of the score matrix
- Examples:
- freescore.cpp.
The documentation for this class was generated from the following file:
- src/scorematrix.hpp