ppALIGN API documentation
freescore.cpp
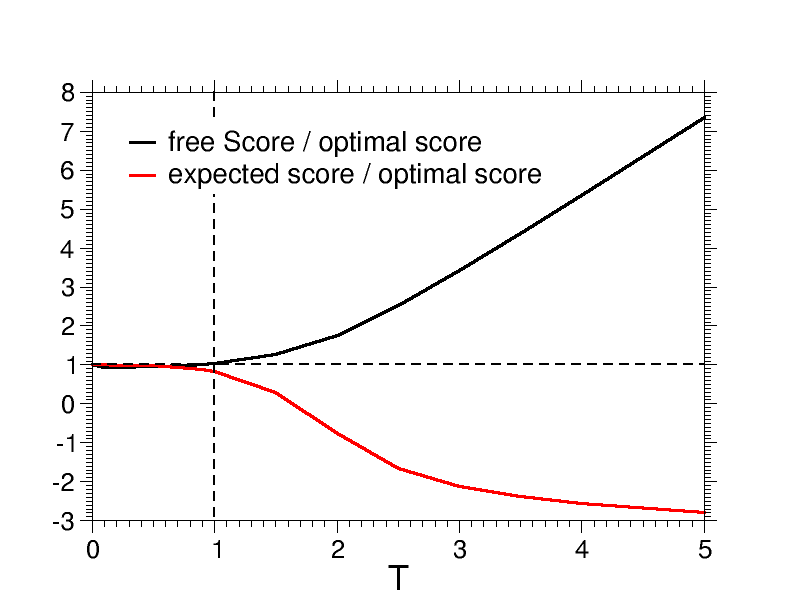
This example shows the usage of the BoltzmannWeight to produce the plot
// Computation of the free and expected score // for the finite temperature model // // There are two methods: #include <partitionworkspace.hpp> #include <weights.hpp> #include <firstmoment.hpp> #include <viterbi.hpp> using namespace std; int main() { // the sequences ProteinSequence seq1("MTLWMHLLPLLPQLPLWGSNLAPEFAEQQGCFSHLNRTVELDVEVEYELCGERT"); ProteinSequence seq2("MELWVRLLPLAPQLALYGPEFIDQHTSIVEALAAYCGGHLGRAICSNLEVEVLYEVCEGRS"); // alignment parameters double temperatures[] = { 0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0, 1.5, 2.0, 2.5, 3.0, 3.5, 4.0, 5.0, 6.0, 7.0, 8.0 }; int n_temper = 20; string matrix_name = "blosum62"; double matrix_scale = 2.0/log(2.0); // the scale of the blosum matrix // only needed when we using the // integer valued matrix int gap_open = 11; int gap_extension = 1; bool allow_DI = true; bool allow_ID = false; // use the integer valued matrix as a basis // this is closer to the optimal alignment based // on the integer score matrix, but there might be // some rounding errors // if we use this option we may expect that // the free_score -> optimal score for small temperatures. bool use_int_matrix = true; // the score matrix ScoreMatrix score_matrix(proteinAlphabet,proteinAlphabet,matrix_name); // the workspace objects PartitionWorkspace pws(10000); ViterbiWorkspace vws(10000); // compute optimal score int score = vws.OptimalScore(seq1, seq2, score_matrix, LocalBounded, gap_open, gap_extension, allow_ID, allow_DI); cout << "#optimal score: " << score << endl; // the scale of the blosum // only needed when we using the // integer valued matrix // estimate lambda from sequence and compare with // sequence composition: Frequencies freq(proteinAlphabet); freq.Count(seq1); freq.Count(seq2); freq.Normalize(); double a = exp(0.0); double lambda_est = score_matrix.GetScale(freq,freq); double lambda = ScoreMatrix::GetScale(proteinAlphabet, proteinAlphabet, matrix_name); cout << "#sequence lambda: " << lambda_est << endl; cout << "#blosum lambda: " << lambda << endl; // the driver for the first moment cout << "#temperature free_score/opt_score expected_score/opt_score" << endl; DriverFirstMoment first_moment(score_matrix,gap_open,gap_extension); for(int i = 0; i < n_temper; i++) { BoltzmannWeight * weight; double temper; // weights for the finite temperature model if(use_int_matrix) { // the canoical temperature is equal to the matrix_scale temper = temperatures[i]*matrix_scale; weight = new BoltzmannWeight(score_matrix, temper, gap_open, gap_extension, LocalBounded, allow_DI, allow_ID); } else { // here, the canonical temperature is 1 temper = temperatures[i]; weight = new BoltzmannWeight(proteinAlphabet, proteinAlphabet, matrix_name, temper, gap_open, gap_extension, LocalBounded, allow_DI, allow_ID); } xdouble Z; double free_score,expected_score; // add driver and perform the computation pws.AddDriver(first_moment); Z = pws.PartitionFunction(seq1, seq2, *weight); // compute free score free_score = log(Z) * temper * weight->scale; // expected score expected_score = first_moment.FirstMoment(); cout << temperatures[i] << "\t" << free_score/score << "\t" << expected_score/score << endl; delete weight; } return 0; }