ppALIGN API documentation
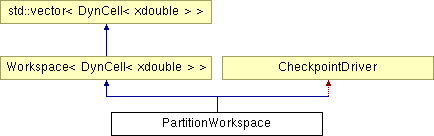
PartitionWorkspace Class Reference
#include <partitionworkspace.hpp>
Classes | |
| struct | ForwardBackwardInfo |
| struct | RowInfo |
| struct | Transition |
| struct | XTransition |
Public Types | |
| typedef Workspace< Transition > ::iterator | trans_iterator |
Public Member Functions | |
| PartitionWorkspace (size_t s) | |
| ~PartitionWorkspace () | |
| void | AddDriver (PartitionDriverBase &driver) |
| void | RemoveDriver (PartitionDriverBase &driver) |
| void | RemoveAllDrivers () |
| xdouble | PartitionFunction (const Sequence &seq1, const Sequence &seq2, const Weight &weight) |
| xdouble | PartitionFunction (const Sequence &seq1, const Sequence &seq2, const Align &a, const Weight &weight, long strip_size=10, long max_strip_size=50, long inc_strip_size=10) |
| Pair | LocalStrip (Sequence &seq1, Sequence &seq2, Align &align, const Weight &weight, long stip_size=10, long max_strip_size=50, long inc_strip_size=10) |
| xdouble | GetNullProb () |
parameters | |
| long | Length1 () const |
| long | Length2 () const |
| Sequence::const_iterator | Seq1Begin () const |
| Sequence::const_iterator | Seq2Begin () const |
| const Weight & | GetWeight () const |
result retrieval | |
| RowInfo & | GetRowInfo (long i) |
| const RowInfo & | GetRowInfo (long i) const |
| RowInfo & | GetLastRowInfo (long i) |
| const RowInfo & | GetLastRowInfo (long i) const |
| const xdouble & | GetZ () const |
| const xdouble & | GetInvZ () const |
| double | GetInsertionProb (long i) const |
| double | GetDeletionProb (long j) const |
| double | GetPairProb1 (long i) const |
| double | GetPairProb2 (long j) const |
Return background probability. | |
| xdouble | GetDanglingForwardInsert (long i) const |
| xdouble | GetDanglingBackwardInsert (long i) const |
| xdouble | GetDanglingForwardDelete (long j) const |
| xdouble | GetDanglingBackwardDelete (long j) const |
Detailed Description
The main computation. This class provides the framework for the main computation of ppALIGN. It performs the forward and backward computation based on the choosen model. Intermediate results (a row of forward and backward quantities) are given to the drivers (of type PartitionDriverBase) that may perform marginalization or sampling.
- Examples:
-
alternative_alignments.cpp, freescore.cpp, and pairprobmap.cpp.
Definition at line 95 of file partitionworkspace.hpp.
Member Typedef Documentation
Iterator over vector of transitions
Definition at line 200 of file partitionworkspace.hpp.
Constructor & Destructor Documentation
| PartitionWorkspace::PartitionWorkspace | ( | size_t | s | ) |
Constructor.
- Parameters:
-
s number of cells that can be used for computation. More cells accelerate the computations.
| PartitionWorkspace::~PartitionWorkspace | ( | ) |
Destructor.
Member Function Documentation
| void PartitionWorkspace::AddDriver | ( | PartitionDriverBase & | driver | ) |
Define a new driver that calculate certain quantities. The driver must tell the main algorithm (class ParitionWorkspace) the required variables
- See also:
- PartitionDriverBase After the computation all drivers are removed and must be added before the next computation
- Parameters:
-
driver an object that performs a certain computation
- Examples:
- alternative_alignments.cpp, and freescore.cpp.
| xdouble PartitionWorkspace::GetDanglingBackwardDelete | ( | long | j | ) | const [inline] |
- Parameters:
-
j index of the second sequence
- Returns:
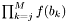
Definition at line 1203 of file partitionworkspace.hpp.
| xdouble PartitionWorkspace::GetDanglingBackwardInsert | ( | long | i | ) | const [inline] |
- Parameters:
-
i index of the first sequence 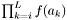
Definition at line 1195 of file partitionworkspace.hpp.
| xdouble PartitionWorkspace::GetDanglingForwardDelete | ( | long | j | ) | const [inline] |
- Parameters:
-
j index of the second sequence
- Returns:
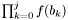 if j >= 0 and 1.0 otherwise
if j >= 0 and 1.0 otherwise
Definition at line 1187 of file partitionworkspace.hpp.
| xdouble PartitionWorkspace::GetDanglingForwardInsert | ( | long | i | ) | const [inline] |
- Parameters:
-
i index of the first sequence
- Returns:
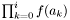 if i >= 0 and 1.0 otherwise
if i >= 0 and 1.0 otherwise
Definition at line 1178 of file partitionworkspace.hpp.
| double PartitionWorkspace::GetDeletionProb | ( | long | j | ) | const [inline] |
Return the marginal probability that b_j is gapped. It is required that there is at least one driver registered that forces the PartitionWorkspace object to perform the backward computation (via PartitionDriverBase::NeedForwardBackward())
- Parameters:
-
j sequence coordinate 0 <= j < Length2()
- Returns:
- Prob(-, b_j)
Definition at line 1015 of file partitionworkspace.hpp.
| double PartitionWorkspace::GetInsertionProb | ( | long | i | ) | const [inline] |
Return the marginal probability that a_i is gapped. It is required that there is at least one driver registered that forces the PartitionWorkspace object to perform the backward computation (via PartitionDriverBase::NeedForwardBackward())
- Parameters:
-
i sequence coordinate 0 <= i < Length1()
- Returns:
- Prob(a_i, -)
Definition at line 1002 of file partitionworkspace.hpp.
| const xdouble & PartitionWorkspace::GetInvZ | ( | ) | const [inline] |
| const PartitionWorkspace::RowInfo & PartitionWorkspace::GetLastRowInfo | ( | long | i | ) | const [inline] |
Get the boundary of the row i in the last iteration of the strip size estimation. If the computation on the full lattice is performed, the return value is row_info.begin_j = 0 and row_info.end_j = Length2().
- Parameters:
-
i row i in sequence coordinates (i=-1 and i=length1 yields the boundary rows)
- Returns:
- RowInfo object
Definition at line 961 of file partitionworkspace.hpp.
| PartitionWorkspace::RowInfo & PartitionWorkspace::GetLastRowInfo | ( | long | i | ) | [inline] |
Get the boundary of the row i in the last iteration of the strip size estimation. If the computation on the full lattice is performed, the return value is row_info.begin_j = 0 and row_info.end_j = Length2().
- Parameters:
-
i row i in sequence coordinates (i=-1 and i=length1 yields the boundary rows)
- Returns:
- RowInfo object
Definition at line 955 of file partitionworkspace.hpp.
| xdouble PartitionWorkspace::GetNullProb | ( | ) | [inline] |
Return the probability that sequences are not locally aligned at all.
- Returns:
- probability
Definition at line 1211 of file partitionworkspace.hpp.
| double PartitionWorkspace::GetPairProb1 | ( | long | i | ) | const [inline] |
Return the marginal probability that a_i is paired It is required that there is at least one driver registered that forces the PartitionWorkspace object to perform the forward-backward computation (via PartitionDriverBase::NeedForwardBackward())
- Parameters:
-
i sequence coordinated 0 <= i < Length1()
- Returns:
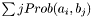
Definition at line 1029 of file partitionworkspace.hpp.
| double PartitionWorkspace::GetPairProb2 | ( | long | j | ) | const [inline] |
Return the marginal probability that b_j is paired. It is required that there is at least one driver registered that forces the PartitionWorkspace object to perform the forward-backward computation (via PartitionDriverBase::NeedForwardBackward()) PartitionDriverBase::NeedBackward())
- Parameters:
-
j sequence coordinated 0 <= j < Length2()
- Returns:
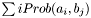
Definition at line 1043 of file partitionworkspace.hpp.
| const PartitionWorkspace::RowInfo & PartitionWorkspace::GetRowInfo | ( | long | i | ) | const [inline] |
Get the boundary of the row i. If the computation on the full lattice is performed, the return value is row_info.begin_j = 0 and row_info.end_j = Length2().
- Parameters:
-
i row i in sequence coordinates (i=-1 and i=length1 yields the boundary rows)
- Returns:
- RowInfo object
Definition at line 948 of file partitionworkspace.hpp.
| PartitionWorkspace::RowInfo & PartitionWorkspace::GetRowInfo | ( | long | i | ) | [inline] |
Get the boundary of the row i. If the computation on the full lattice is performed, the return value is row_info.begin_j = 0 and row_info.end_j = Length2().
- Parameters:
-
i row i in sequence coordinates (i=-1 and i=length1 yields the boundary rows)
- Returns:
- RowInfo object
Definition at line 942 of file partitionworkspace.hpp.
| const Weight & PartitionWorkspace::GetWeight | ( | ) | const [inline] |
Model paramters.
- Returns:
- a constant reference to the model parameters.
Definition at line 997 of file partitionworkspace.hpp.
| const xdouble & PartitionWorkspace::GetZ | ( | ) | const [inline] |
| long PartitionWorkspace::Length1 | ( | ) | const [inline] |
Length of the first sequence.
- Returns:
- the length of the first sequence that has been given in the methods PartitionFunction()
Definition at line 975 of file partitionworkspace.hpp.
| long PartitionWorkspace::Length2 | ( | ) | const [inline] |
Length of the second sequence.
- Returns:
- the length of the second sequence that has been given in the methods PartitionFunction()
Definition at line 980 of file partitionworkspace.hpp.
| Pair PartitionWorkspace::LocalStrip | ( | Sequence & | seq1, | |
| Sequence & | seq2, | |||
| Align & | align, | |||
| const Weight & | weight, | |||
| long | stip_size = 10, |
|||
| long | max_strip_size = 50, |
|||
| long | inc_strip_size = 10 | |||
| ) |
Reduce alignment to strip around local alignment.
- Parameters:
-
[in,out] seq1 [in,out] seq2 [in,out] align
| xdouble PartitionWorkspace::PartitionFunction | ( | const Sequence & | seq1, | |
| const Sequence & | seq2, | |||
| const Align & | a, | |||
| const Weight & | weight, | |||
| long | strip_size = 10, |
|||
| long | max_strip_size = 50, |
|||
| long | inc_strip_size = 10 | |||
| ) |
Compute the forward and / or backward probabilities around the alignment a.
- Parameters:
-
seq1 input sequence seq2 input sequence a alignment around which the computation is performed weight weights definition of the model strip_size inital size of the strip max_strip_size max. size of the strip if the required strip size exceeds this number the computation is performed on the complete space. inc_strip_size increase the strip by this number when estimating the strip size
| xdouble PartitionWorkspace::PartitionFunction | ( | const Sequence & | seq1, | |
| const Sequence & | seq2, | |||
| const Weight & | weight | |||
| ) |
Compute the forward and / or backward probabilities. The computation is performed on the complete quadratic search space.
- Parameters:
-
seq1 first input sequence seq2 second input sequence weight definition of the model.
- Examples:
- alternative_alignments.cpp, and freescore.cpp.
| void PartitionWorkspace::RemoveAllDrivers | ( | ) |
Remove all registered drivers. If the workspace is to be reused for further computation the drivers have to be add again via AddDriver.
| void PartitionWorkspace::RemoveDriver | ( | PartitionDriverBase & | driver | ) |
Remove a driver and corresponding callback functions
| Sequence::const_iterator PartitionWorkspace::Seq1Begin | ( | ) | const [inline] |
Iterator to the begin of the first sequence.
- Returns:
- an iterator that points to the begin of the first sequence given in the methods PartitionFunction()
Definition at line 986 of file partitionworkspace.hpp.
| Sequence::const_iterator PartitionWorkspace::Seq2Begin | ( | ) | const [inline] |
Iterator to the begin of the second sequence.
- Returns:
- an iterator that points to the begin of the second sequence given in the methods PartitionFunction()
Definition at line 991 of file partitionworkspace.hpp.
The documentation for this class was generated from the following file: