ppALIGN API documentation
ViterbiWorkspace Class Reference
#include <viterbi.hpp>
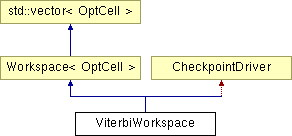
Inheritance diagram for ViterbiWorkspace:
Public Member Functions | |
| ViterbiWorkspace (size_t n) | |
| int | OptimalAlignment (const Sequence &seq1, const Sequence &seq2, const ScoreMatrix &mat, AlignmentType atype, int gap_open, int gap_extension, bool _allow_DI, bool _allow_ID, Align &a, bool _dangling=true) |
| int | OptimalScore (const Sequence &seq1, const Sequence &seq2, const ScoreMatrix &mat, AlignmentType atype, int gap_open, int gap_extension, bool _allow_DI, bool _allow_ID) |
Detailed Description
Workspace for optimal alignments (local or global)
- Examples:
Definition at line 48 of file viterbi.hpp.
Constructor & Destructor Documentation
| ViterbiWorkspace::ViterbiWorkspace | ( | size_t | n | ) |
Constructor.
- Parameters:
-
n number of dynamic programming cells to be used.
Member Function Documentation
| int ViterbiWorkspace::OptimalAlignment | ( | const Sequence & | seq1, | |
| const Sequence & | seq2, | |||
| const ScoreMatrix & | mat, | |||
| AlignmentType | atype, | |||
| int | gap_open, | |||
| int | gap_extension, | |||
| bool | _allow_DI, | |||
| bool | _allow_ID, | |||
| Align & | a, | |||
| bool | _dangling = true | |||
| ) |
Determine optimal score and alignment. Determine optimal alignment of the sequences seq1 and seq2 by SmithWaterman (atype == LocalBounded) or Needleman-Wunsch (atype == Global or atype == GlobalBounded) algorithm.
- Parameters:
-
seq1 first input sequence seq2 second input sequence mat score matrix atype alignment type gap_open gap open parameter gap_extension gap extension parameter allow_ID allow a deletion followed by insertion allow_DI allow an insertion followed by deletion a output alignment _dangling if true fill begin and end of local alignment with gaps (only relevant for atype == LocalBounded)
- Returns:
- optimal score
The documentation for this class was generated from the following file:
- src/viterbi.hpp